Contents
Overview
The ![]() Find Group Specific SNVs function can be used to identify SNVs that are unique to a group of Samples (target-genomes), and do not
appear in a second group of Samples (nontarget-genomes). The function is available in the comparison table, and extracts the SNVs from the allele sequences that are corresponding to the distance columns (those with green colored headings) of the comparison table.
Find Group Specific SNVs function can be used to identify SNVs that are unique to a group of Samples (target-genomes), and do not
appear in a second group of Samples (nontarget-genomes). The function is available in the comparison table, and extracts the SNVs from the allele sequences that are corresponding to the distance columns (those with green colored headings) of the comparison table.
![]() Hint: The AMRFinderPlus Task Template does not support currently this function.
Hint: The AMRFinderPlus Task Template does not support currently this function.
Starting Group Specific SNV Search
When the Find Group Specific SNVs function is invoked, a handling of missing values dialog window is shown if the comparison table contains any distance columns with missing data. By default all Samples with more than 10% missing targets will be removed. Alternatively or in addition all columns with any missing data can be removed. If no missing data exist at all, this dialog is skipped and the following dialog is shown only.
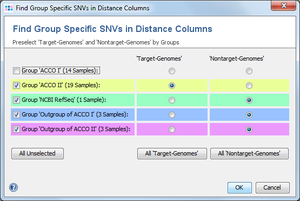
If color groups are defined in the comparison table, a group selection dialog is shown that allows to allocate each color group to the target-genomes, nontarget-genomes, or exclusion-genomes group. If no color groups exist in the comparison table, this dialog is skipped and the following dialog is shown only.
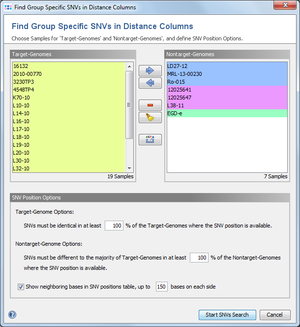
The following Sample selection and options dialog is always shown. If color groups are defined and thus a group selection has been made in the previous dialog all Samples are already allocated either to the target-genomes or nontarget-genomes groups. If corrections are needed the Samples can still be removed or moved here between the two groups. If no group selection has been done previously all Samples are initially part of the target-genomes group and therefore need to be moved accordingly into the two groups.
The export button ![]() allows to write the FASTA files of all Samples into two different directories (target-genomes and nontarget-genomes). These assembly files can next be used for a search of a stretch of unique bases, i.e., a signature, with external software (e.g., PanSeq Novel Region Finder).
allows to write the FASTA files of all Samples into two different directories (target-genomes and nontarget-genomes). These assembly files can next be used for a search of a stretch of unique bases, i.e., a signature, with external software (e.g., PanSeq Novel Region Finder).
Below the Sample selection section the SNV Position Options can be specified:
- Target-Genomes Option:
- The minimum percentage of target-genome Samples, that must be identical in the SNV position. Samples that don't have a allele at all (because it was not found in the scan procedure) are not counted here. This threshold is set by default to 100%.
- Nontarget-Genomes Option:
- The minimum percentage of nontarget-genome Samples, that must be different in the SNV position to the majority of target-genomes in this position. Samples that don't have a allele at all (because it was not found in the scan procedure) are not counted here. This threshold is set by default to 100%.
- Show neighboring bases in SNV positions table
- If this option is enabled, the resulting table contains additional columns with neighboring bases next to the SNV position. The export of neighboring bases can be used to design primers (e.g., to confirm by Sanger sequencing the SNVs). A column with neighboring bases contains up to a specified number of bases (default 150) from each side of the SNV position. The SNV position itself is indicated with a lowercase character. This column is followed by a second column with the number of bases that were exported (default 301 or lower, if the SNV position lies near the start or end of a target), and a third column with the SNV position. There are three different types of neighboring bases: exported from the first target-genome Sample that contains this SNV, the ref.-seq. of the target, and the seed genome (if applicable also with intergenic region bases) of the Task Template. If the target or the SNV is not found in the first target-genome Sample, additional columns from target-genome Samples that contain the SNV are inserted next to the first target-genome Sample columns.
Result of Group Specific SNV Search
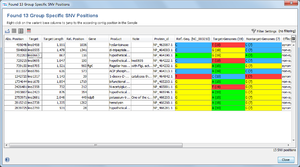
The result of the group specific SNV search is shown in a SNV-centric table, where each row represents an group specific SNV position that was found. The columns Target-Genomes and Nontarget-Genomes summarizes the bases appearing in the according set of Samples, together with their frequencies. By selecting a SNV row and then right-clicking the target sequence can be exported.
The following functions are available via the toolbar:
- Export Table: Export the table to an Excel or CSV file.
- Multiple Alignment: Once a row is selected this command creates a multiple alignment for the target of the select SNV position for some or all Samples. It is not recommended to do a multiple alignment of more than 100 Samples due to time and especially memory constrains. Therefore, a Sample selection dialog is shown first that allows also to remove Samples from multiple alignment analysis.
- Show/Hide all Sample SNV Columns: By default only the summary columns for the variant bases is shown. Use this function to show all Sample variant base columns.
- Open in Comparison Table: The function can be used to create a Sample-centric comparison table that allows to calculate and visualize distances between Samples based on the nucleotides of the found SNV positions.
- Filter Settings: The Filter SNV Positions dialog can be used to show only a subset of the SNV positions in the table. Up to two different filter options can be specified:
- Show only substitution SNV positions (hide insertions/deletions)
- This hides all SNV positions that have any insertions or deletions. This option is by default turned on and only shown if there are at all any insertions or deletions present.
- Show only SNV positions where the majority target-genome SNV is synonymous in comparison to Ref.-Seq.
- This hides all SNV positions that have an non-synonymous change in the amino acid of the majority target-genome SNV (compared to Ref.-Seq. column). The synonym/non-synonym effect is also shown in the column Effect to Ref.-Seq.
- Show only SNV positions that change the GC content
- This hides all SNV positions that do not increase or decrease the GC content (i.e., A<->T and G<->C) between all target-genomes and all nontarget-genomes. This option is useful when the intention is to design high-resolution melting curve (HRMC) assay.
Using Group Specific SNVs for Screening Assays
The general purpose of the Find Group Specific SNVs function is to identify target-genome group specific, i.e., unique, SNVs that can be used to develop highly specific screening assays (e.g., High Resolution Melting Curve [HRMC], Melt-MAMA [PubMed 22438886], or TaqMan™ realtime PCR). For doing so usually it is beneficial to export the SNV table and the target sequences.
Once primers and/or probes have been designed (e.g., using the ABI Primer Express tool) the specificity of those oligonucleotides can be checked via the above mentioned Multiple Alignment functionality. Thus, select the target row for which primer/probes were designed and press the Multiple Alignment button. Then select all or a subset of the Samples and confirm the upcoming dialog.
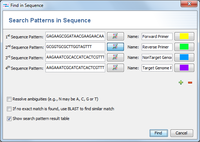
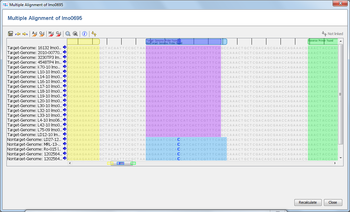
The Samples are now loaded and a multiple alignment is calculated for the alleles (this can take a while if a large number of Samples were selected). Finally, the resulting multiple alignment is opened in a new window. Press the ![]() Find button above the sequence alignment, and enter the bases of the primer/probes that were designed (optionally enter also a name). Each primer can be highlighted with a different color. By default the option Show search pattern result table is checked. Once the dialog is confirmed, the matches of the primer/probes are highlighted in the multiple alignment and a search pattern result table opens if the according option was chosen. Patterns with multiple hits are not reported in the search pattern result table.
Find button above the sequence alignment, and enter the bases of the primer/probes that were designed (optionally enter also a name). Each primer can be highlighted with a different color. By default the option Show search pattern result table is checked. Once the dialog is confirmed, the matches of the primer/probes are highlighted in the multiple alignment and a search pattern result table opens if the according option was chosen. Patterns with multiple hits are not reported in the search pattern result table.